An independent laboratory
Nanoparticle innovation in search of a cancer cure
Celebrating 36 years of innovation!
All profits support our research.
Learn more about our:
- Nanoparticle Therapies: Our Cancer Research
- Nanoparticle Technology for your Lab:
Nanoparticle Products for Imaging, Microscopy and Biodetection
Home of the original Nanogold®
Our mission at Nanoprobes is to harness chemistry, biology and nanotechnology to engineer the most sensitive reagents and nanotechnology for detecting, locating and visualizing biological molecules.
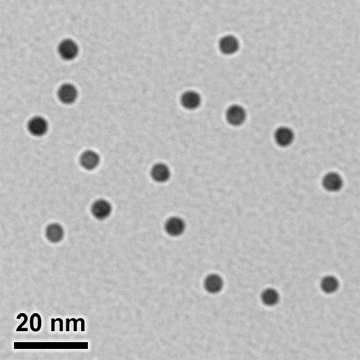
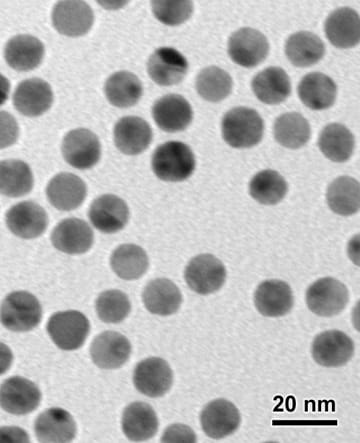
Our unique gold nanoparticle labeling technology uses chemically cross-linked metal clusters and nanoparticles as labels.
Unlike conventional immunogold probes, in which colloidal gold particles are electrostatically adsorbed to antibodies and proteins, our gold nanoparticle labels are uncharged molecules which are cross-linked to specific sites on biomolecules. This gives our nanogold probes greater range and versatility than colloidal gold.
Our labels can be attached to any molecule with a reactive group — proteins, peptides, oligonucleotides, small molecules and lipids — for detection and localization. Other labels can be combined with our gold particle labels; our unique FluoroNanogold probes combine Nanogold® and fluorescein into a single probe for imaging a specimen both by fluorescence and electron microscopy. Our gold nanoparticles, including our Nanogold™ conjugates, and Nanogold™ labeling reagents, are formulated for use with electron microscopy (tem) as well as light microscopy, immunolabeling, x-ray contrast for EM, angiography, mammography and microCT, and for correlative and fluorescence microscopy imaging.
New probes can be engineered based on any fragment of a naturally occurring biomolecule, and the label can be positioned away from the binding site so it does not interfere with binding.
Our Nanogold® probes have been cited in over 250 publications, and are used in the laboratories of several Nobel Laureates.