Nanogold in cryo-electron tomography:
Determining receptor structure and function
The neonatal Fc receptor (FcRn) is part of the system that gives immunity to the fetus or newborn until its own immune system develops. It transports maternal IgG across epithelial barriers: in newborn rats, it transfers IgG from milk to blood by apical-to-basolateral transcytosis across intestinal epithelial cells. This is driven by pH: FcRn binds IgG at the lower, apical pH (6.0 - 6.5), then releases it at the higher basolateral pH (7.4). Maternal IgG from milk is taken up by FcRn-expressing cells in the proximal small intestine (duodenum and jejunum).
Pamela Bjorkman and Wanzhong He of the CalTech Biology Department used electron tomography with Nanogold®-labeled Fc to visualize jejunal transcytosis directly in space and time: to do so, they developed a new labeling and detection method to map individual Nanogold®-labeled Fc within transport vesicles, while simultaneously characterizing the vesicles by immunolabeling. In contrast, a 1 nm colloidal gold probe aggregated and could not access the target, and even a 3 nm covalently functionalized gold particle did not show target binding.
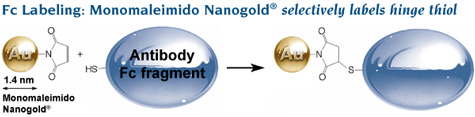
Labeling of Fc using Monomaleimido Nanogold® to label a hinge thiol. Comparison of probe size and resolution: conventional colloidal gold IgG is much larger than Nanogold®-Fc and provides much lower resolution. |
Fc was selectively labeled with Monomaleimido Nanogold® at a hinge thiol site, where it does not interfere with FcRn binding. Fc was reduced with mercaptoethylamine hydrochloride (MEA), incubated for 24 hours with reconstituted Monomaleimido Nanogold®, and conjugates isolated by gel filtration (Superdex 75 column) then FcRn affinity chromatography. Spectrophotometric calculation gave 0.8 Nanogold® per Fc, and gel filtration confirmed that the conjugates were monomeric.
To investigate how it works in the body, Au-Fc was fed to neonatal rats, and the small intestine was harvested. Intestinal segments were processed either (a) by chemical fixation followed by gold enhancement, OsO4 and uranyl acetate, or (b) by high-pressure freezing/freeze substitution fixation (HPF/FSF).
Because conventional pre-embedding enhancement protocols are incompatible with freeze-substitution fixation, the authors developed a special silver enhancement method conducted entirely in acetone:5 saturated solutions of silver nitrate (0.04%), hydroquinone (0.3%), citric acid (0.4%) and sucrose (0.1%), in pure acetone were cooled to -20°C overnight in the dark, centrifuged, then cooled to 50°C and mixed in pre-cooled containers for use.
They enlarged the Nanogold® to at most 8 nm using this method, then applied gold toning to protect it from osmium etching, then enlarged to 10-16 nm with gold enhancement. To check how the FcRn labeling related to the endocytosis, immunolabeling for early, late and recycling endosomal markers was performed on some of the gold-enhanced intestinal cryosections using a modified Tokuyasu method for on-section immunolabeling.
Labeled samples were infiltrated with resin and polymerized. EM tomography was conducted on 120 - 200 nm sections using a FEI T12 transmission electron microscope operating at 120 kV. Dual-axis tilt series were collected at x 6,500 at 700 nm underfocus, using enlarged golds as alignment markers, then combined to form a dual-axis tomogram and reconstructed by manual segmentation using the IMOD software package.
The results showed a complex system of vesicle transport and association, in which the labeled Fc migrates to the basolateral surface through a network of tubular and interconnected vesicles. Markers for early, late and recycling endosomes labeled vesicles with different but overlapping morphologies, showing that FcRn trafficking is complex.
However, localization of native FcRn did not reveal how it interacted with vesicle membranes at the molecular level. Therefore, the group investigated the molecular interaction of the Au-Fc with a synthetic liposomal preparation of recombinant soluble FcRn. This time, they used electron cryotomography to localize the Nanogold® in 250 nm frozen-hydrated sections, with 10 nm colloidal gold added as fiducial markers for alignment.
Molecular interaction of Au-Fc with liposomes.
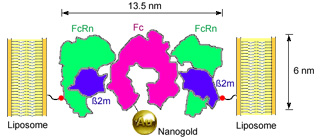
Binding of Nanogold®-labeled Fc to FcRn-liposomes. Residue 19 of the FcRn light chain (beta-2-microglobulin), which was mutated to cysteine in the K19C FcRn mutant, is highlighted as a red sphere linked to the liposome surface. |
Cryo-electron microscopy (cryo-EM), in which ‘fixation’ is accomplished through freezing, is one of the best ways there is to look at membranes and vesicles: the aqueous compartments are preserved, and fixatives and other chemicals that can change the morphology of the specimens are avoided, so lipid bilayers are preserved much closer to their natural state.
Specimens were loaded on glow-discharged Lacey carbon grids and plunge-frozen in liquid ethane at liquid nitrogen temperature. Tilt series for tomographic reconstructions (-60° to 60°; 1.5° step size) were collected on a FEI Tecnai F30 microscope at 300 kV with defocus set to 4.5–8.0 mm at 0°. As before, IMOD was used for 3D reconstructions. With unlabeled Fc, a strip of higher density was observed between adjoining FcRn-liposomes, indicating FcRn-Fc complexes that bridged between adjacent liposome; however, with Nanogold®-labeled Fc, individual FcRn-Fc complexes could be located by the Nanogold® particle densities.
Localization of Fc receptor with Nanogold®
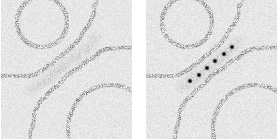
Schematic showing EM localization of (Left) electron density using unlabeled Fc and (Right) Nanogold® particles using Nanogold®-Fc. |
Individual Nanogold® labels were clearly resolved in tomograms recorded from 4.5 to 8 mm defocus; so were protein densities, and both leaflets of the liposome bilayer. Nanogold® proved to be a highly effective molecular label, even in 250 nm thick ice, and even though the tomograms were low-pass filtered to the first zero of the contrast transfer function. Therefore, Nanogold® a useful tool to identify single molecules under frozen hydrated conditions, and should work well in systems such as virus-receptor or cell-cell interactions.
Nanogold in cryo-electron tomography:
Visualizing protein structures in intact bacteria
Because it is covalently linked, you can – if you have high enough resolution – localize Nanogold® to the specific region on a protein to which it is attached. Cryo-electron tomography lets you do it in whole organisms, and this is illustrated by Amat, Downing and group in their analysis of the surface layer protein of the bacterium Caulobacter crescentus.
Previously, this protein had been studied only by negative staining of detached surface layer fragments, but this is of limited value because it does not reveal the extent to which this unusually crystalline protein retains long-range order, or how it relates to the rest of the bacterium. With cryotomography, the protein could be visualized in the intact bacterium.
The authors labeled genetically modified S-layer protein (RsaA) using Monomaleimido Nanogold® at four different engineered cysteine sites: Cys2, Cys277, Cys353, and Cys944. Their microscopic procedure was very similar to that of Bjorkman and He: aliquots of labeled bacteria were taken directly from the culture solution, applied to glow-discharged lacey carbon grids pre-treated with 10 nm gold fiducials, then plunge-frozen in liquid ethane. Images were acquired on a JEOL3100 electron microscope operating at 300 kV, using a cryo-transfer stage cooled to 80K with liquid nitrogen. Tilt series (typically 62° to 62° ±2° with increments of 1° or 2°) were acquired under low-dose conditions and reconstructed with IMOD.
This group, however, had several factors in their favor for even higher resolution. The RsaA protein forms a regular array with six-fold symmetry, and the authors used subtomogram averaging, assisted by symmetry constraints, to reveal Nanogold® electron density which could not be resolved in the individual protein units, and to compensate for missing subunits.
As a result, Nanogold® could be distinguished at each of the labeling sites through its distinct density pattern, and this was used to locate the N- and C-terminals of the protein within the lattice. The authors concluded that Nanogold® could be localized to specific 200 – 300 amino acid residue regions – much higher resolution than is possible with antibody labeling.
- Chen, X.; Winters, C. A., and Reese, T. S.: Life inside a thin section: tomography. J. Neurosci., 28, 9321-9327 (2008).
- Sousa, A. A.; Aronova, M. A.; Kim, Y. C.; Dorward, L. M.; Zhang, G, and Leapman, R. D.: On the feasibility of visualizing ultrasmall gold labels in biological specimens by STEM tomography. J. Struct. Biol., 159, 507-522 (2007).
- He, W.; Ladinsky, M. S.; Huey-Tubman, K. E.; Jensen, G. J.; McIntosh, J. R., and Björkman P. J.: FcRn-mediated antibody transport across epithelial cells revealed by electron tomography. Nature, 455, 542-546 (2008).
- He, W.; Jensen, G. J., and Björkman, P. J.: Nanogold as a Specific Marker for Electron Cryotomography. Microsc. Microanal., 15, 183-188 (2009).
- Abstract from Cambridge University Press
- Morphew, M.; He, W.; Bjorkman, P. J., and McIntosh, J. R.: Silver enhancement of Nanogold particles during freeze substitution for electron microscopy. J. Microsc., 230, 263-267 (2008).
- Amat, F.; Comolli, L. R.; Nomellini, J. F.; Moussavi, F.; Downing, K. H.; Smit, J., and Horowitz, M.: Analysis of the intact surface layer of Caulobacter crescentus by cryo-electron tomography. J. Bacteriol., 192, 5855-5865 (2010).
|
Also in this issue: |